Last time I outlined the derivation of a Metropolis-Hastings algorithm for RJMCMC and this time I want to implement it in Stata for a very simple problem. I have taken the problem from the paper in The American Statistician by Barker and Link that I referenced at the end of my last posting.
We have two binomial samples y1=8, n1=20 and y2=16, n2=30, but we are not sure whether to model them as,
M1: binomials with different probabilities π1 and π2
M2: binomials with a common probability ξ.
We will need priors. In their paper, Barker and Link use identical vague priors that simply cancel out of the equations, so you cannot see exactly what is going on. I prefer to set different priors and I’ll arbitrarily choose, p(M1)=0.3 and p(M2)=0.7. Under M1 my priors on the parameters will be p(π1|M1)=Beta(5,5) and p(π2|M1)=Beta(10,10) and under M2, p(ξ|M2)=Beta(20,20).
Before setting up the RJMCMC program let us analyse the two models separately.
Here is a standard MH program for analysing model M1. It uses the truncated MH sampler, mhstrnc, to ensure that all proposals for π1 and π2 are between 0 and 1. I have lazily used the function, logdensity; this is not very efficient, but the program is so simple that speed is not as issue.
program logpost
args lnf b
local pi1 = `b'[1,1]
local pi2 = `b'[1,2]
scalar `lnf’ = 0
logdensity binomial `lnf’ 8 20 `pi1′
logdensity binomial `lnf’ 16 30 `pi2′
logdensity beta `lnf’ `pi1′ 5 5
logdensity beta `lnf’ `pi2′ 10 10
end
matrix b = (0.5, 0.5)
mcmcrun logpost b using temp.csv , replace ///
param(pi1 pi2) burn(500) update(5000) ///
samplers( 2(mhstrnc , sd(0.5) lb(0) ub(1) ) )
insheet using temp.csv, clear
mcmcstats *
Here is a summary of the chain,
------------------------------------------------------- Parameter n mean sd sem median 95% CrI ------------------------------------------------------ pi1 5000 0.431 0.088 0.0030 0.428 ( 0.260, 0.615 ) pi2 5000 0.521 0.071 0.0022 0.522 ( 0.376, 0.656 ) -------------------------------------------------------
Mixing is fine so I will not bother with the graphics.
Probably you know that the beta prior is conjugate to the binomial probability so it can be shown theoretically that the posterior will also have a beta distribution. When the data are y out of n and the prior on p is Beta(a,b) then the posterior on p will be Beta(a+y,b+n-y).
We could take advantage of this and program our own Gibbs sampler. This new algorithm ought to converge quicker than the Metropolis-Hastings algorithm and be faster too.
program myGibbs
args logpost b ipar
matrix `b'[1,1] = rbeta(13,17)
matrix `b'[1,2] = rbeta(26,24)
end
matrix b = (0.5, 0.5)
mcmcrun logpost b using temp.csv , replace ///
param(pi1 pi2) burn(500) update(5000) ///
samplers( myGibbs , dim(2) )
insheet using temp.csv, clear
mcmcstats *
Now the results are,
------------------------------------------------------- Parameter n mean sd sem median 95% CrI ------------------------------------------------------ pi1 5000 0.433 0.088 0.0012 0.431 ( 0.270, 0.612 ) pi2 5000 0.520 0.070 0.0010 0.522 ( 0.380, 0.654 ) -------------------------------------------------------
As expected we have the same answer but we have reduced the sampling error as measured by the MCMC standard error (sem).
We can even calculate the true posterior mean since Wikipedia tells us that a Beta(a,b) distribution has a mean a/(a+b), so a Beta(13,17) should have a mean 0.433 and a Beta(26,24) should have a mean 0.520, just as we found.
We can do the same for model 2 although I will not present all of the details as they are so similar. If y1=8, n1=20 and y2=16, n2=30 have a common binomial probability with prior, Beta(20,20), the posterior will be Beta(8+16+20,12+14+20)=Beta(44,46) so the posterior mean is 0.489.
Now we will set up a chain that moves between the two models. When it is in M1 the chain will estimate (π1, π2) and should find exactly the same posterior that we have already derived. When the chain is in model 2 it should estimate ξ and again get the same estimate as before. The extra information will come about because the algorithm will decide whether to spend its time in M1 or M2 and it will do so in such a way that the time will be proportional to the model’s posterior probability.
Model M1 has 2 parameters and model M2 has 1 parameter. So in the notation that I set up last week, the maximal set of parameters is, Ψ = (Ψ1, Ψ2) and depending on the choice of model, we need to convert (Ψ1, Ψ2) into (π1, π2) or (Ψ1, Ψ2) into (ξ,u) where u is an arbitrary, unused parameter.
Now we need to know how to move between models.
There is no unique way to jump between the models, but as long as the jumps are 1:1 and we do not enter any cul-de-sacs, we can do what we like, although daft choices will lead to poor mixing and a very long chain.
For instance, if under M1 we currently have estimates (0.5,0.3), it would be odd to propose a move to M2 with ξ =0.9. It would mean that we think that the separate probabilities are 0.5 and 0.3 and we try a move to a model with a common probability that is almost 1. Obviously such a move would rarely be accepted and the chain would take a long time to converge. However, it would eventually move and it would eventually recover; though eventually might mean millions of iterations.
I’m going to start with the simplest possible proposal,
M1: π1 = Ψ1 and π2= Ψ2
M2: ξ = Ψ1 and u=Ψ2
Last time I derived the form of the Metropolis-Hastings algorithm and mentioned that we need to be aware of the problem of parameter transformation and the way that transformation affects the increments dΨ in the Metropolis-Hastings acceptance probability. Here both of the sets of parameters are derived, untransformed, from Ψ. Depending on how you choose to think about it, there is either no transformation of parameters or the transformation is based on simple equality, so either the dΨ cancel directly, or both of the Jacobians are equal to 1 and can be omitted.
This leads us to our first RJMCMC program
program myRJMCMC
args logpost b ipar
local M = `b'[1,1]
if( `M’ == 1 ) {
* currently in model 1: update
matrix `b'[1,2] = rbeta(13,17)
matrix `b'[1,3] = rbeta(26,24)
local psi1 = `b'[1,2]
local psi2 = `b'[1,3]
* try switch to M=2
local a1 = log(binomialp(20,8,`psi1′))+log(binomialp(30,16,`psi2′))+ ///
log(betaden(5,5,`psi1′))+log(betaden(10,10,`psi2′))+log(0.3)
local a2 = log(binomialp(20,8,`psi1′))+log(binomialp(30,16,`psi1′))+ ///
log(betaden(20,20,`psi1′))+log(betaden(26,24,`psi2′))+log(0.7)
if log(runiform()) < `a2′ – `a1′ matrix `b'[1,1] = 2
}
else {
* currently in model 2: update
matrix `b'[1,2] = rbeta(44,46)
matrix `b'[1,3] = rbeta(26,24)
local psi1 = `b'[1,2]
local psi2 = `b'[1,3]
* try switch to M=1
local a1 = log(binomialp(20,8,`psi1′))+log(binomialp(30,16,`psi2′))+ ///
log(betaden(5,5,`psi1′))+log(betaden(10,10,`psi2′))+log(0.3)
local a2 = log(binomialp(20,8,`psi1′))+log(binomialp(30,16,`psi1′))+ ///
log(betaden(20,20,`psi1′))+log(betaden(26,24,`psi2′))+log(0.7)
if log(runiform()) < `a1′ – `a2′ matrix `b'[1,1] = 1
}
end
matrix b = (1, 0.5, 0.5)
mcmcrun logpost b using temp.csv , replace ///
param(m psi1 psi2) burn(500) update(5000) ///
samplers( myRJMCMC , dim(3) )
insheet using temp.csv, clear
tabulate m
mcmcstats *
mcmcstats * if m == 1
mcmcstats * if m == 2
The vector of parameters, b, has been expanded to contain an indicator of the model as well as Ψ.
Let’s note a few of the features of the program.
(a) I have programmed for clarity rather than for efficiency, for instance, I copy the parameter values into locals which strictly I do not need to do.
(b) First I find out which model the chain is currently in. Then I update the parameter estimates just as I did when I was analysing the models separately. Finally I decide whether to stay with the current model or to switch and I do this with a Metropolis-Hastings step.
(c) I calculate the log of the acceptance ratio because I want to avoid loss of precision due to very small probabilities.
(d) Each part of the acceptance ratio involves the likelihood, the prior on Ψ for that model and the prior on the model. Often there would also be a Jacobian but as noted already, the Jacobians for our moves are both 1.
(e) The proposal probabilities for the moves (g() in the notation that I set up last time) are omitted from the acceptance ratio because they cancel. Since there are only two models, we propose a move to M2 when in M1 and a move to M1 when in M2 with equal probability.
(f) Under model 2 I still have to update u even though it does not occur in the model’s likelihood. Here I have chosen to use the same distribution that I used for π2 under model 1, but this is only for convenience. The choice is arbitrary.
Here are the results
m | Freq. Percent Cum. ------------+----------------------------------- 1 | 1,165 23.30 23.30 2 | 3,835 76.70 100.00 ------------+----------------------------------- Total | 5,000 100.00
So we spend about 77% of the time in model 2, which means that the posterior probability of model 2 is 0.77. Our prior was 0.7 so the data have pushed us slightly further in the direction of model 2.
We can even calculate the Bayes Factor. The prior ratio of model probabilities is 0.7/0.3=2.33 and the posterior ratio is 0.77/0.23=3.35 so the Bayes Factor is 3.35/2.33=1.44. Not a very impressive Bayes Factor since it suggests that the data only push us weakly in the direction of Model 2; most of our preference for model 2 comes from the prior.
The Stata command
. count if m != m[_n-1]
Tells us that we switched model 1785 times in the chain of 5,000 iterations. The ideal for MH algorithms is to move around 30%-50% of the time so this represents good mixing.
If we extract the results for the time that the chain spent in Model 1
---------------------------------------------------------------------------- Parameter n mean sd sem median 95% CrI ---------------------------------------------------------------------------- m 1165 1.000 0.000 . 1.000 ( 1.000, 1.000 ) psi1 1165 0.435 0.091 0.0028 0.436 ( 0.262, 0.618 ) psi2 1165 0.519 0.073 0.0021 0.518 ( 0.371, 0.657 ) ----------------------------------------------------------------------------
The parameter estimates are very similar to the values that we calculated before (0.433 and 0.520).
Extracting the values for Model 2 we get
---------------------------------------------------------------------------- Parameter n mean sd sem median 95% CrI ---------------------------------------------------------------------------- m 3835 2.000 0.000 . 2.000 ( 2.000, 2.000 ) psi1 3835 0.489 0.052 0.0009 0.490 ( 0.387, 0.591 ) psi2 3835 0.521 0.070 0.0011 0.523 ( 0.379, 0.654 ) ----------------------------------------------------------------------------
Once again the estimate of the joint probability is very close to the theoretical mean (0.489). The second parameter is the u and must be ignored.
Basically that is RJMCMC, but let us try some variations.
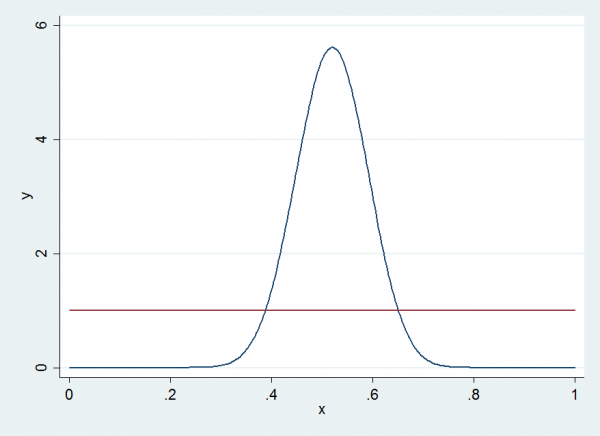
First I said that the updating of u is arbitrary, so I will change from Beta(26,24) to Beta(1,1), which is equivalent to using a uniform distribution. Here is the density of the distribution that I used before and the uniform that I will switch to; they look very different but since that will be used for u, they may affect the mixing but should not change the posteriors.
The code is almost the same. I’ve highlighted the changes in red.
program myRJMCMC
args logpost b ipar
local M = `b'[1,1]
if( `M’ == 1 ) {
matrix `b'[1,2] = rbeta(13,17)
matrix `b'[1,3] = rbeta(26,24)
local psi1 = `b'[1,2]
local psi2 = `b'[1,3]
* try switch to M=2
local a1 = log(binomialp(20,8,`psi1′))+log(binomialp(30,16,`psi2′))+ ///
log(betaden(5,5,`psi1′))+log(betaden(10,10,`psi2′))+log(0.3)
local a2 = log(binomialp(20,8,`psi1′))+log(binomialp(30,16,`psi1′))+ ///
log(betaden(20,20,`psi1′))+log(betaden(1,1,`psi2′))+log(0.7)
if log(runiform()) < `a2′ – `a1′ matrix `b'[1,1] = 2
}
else {
matrix `b'[1,2] = rbeta(44,46)
matrix `b'[1,3] = rbeta(1,1)
local psi1 = `b'[1,2]
local psi2 = `b'[1,3]
* try switch to M=1
local a1 = log(binomialp(20,8,`psi1′))+log(binomialp(30,16,`psi2′))+ ///
log(betaden(5,5,`psi1′))+log(betaden(10,10,`psi2′))+log(0.3)
local a2 = log(binomialp(20,8,`psi1′))+log(binomialp(30,16,`psi1′))+ ///
log(betaden(20,20,`psi1′))+log(betaden(1,1,`psi2′))+log(0.7)
if log(runiform()) < `a1′ – `a2′ matrix `b'[1,1] = 1
}
end
The results are so similar that I will not even bother to show them. The mixing is slightly worse with a tendency to stay longer in the same model before switching, but even that effect is minor.
At present the algorithm always updates the parameters for the current model and then always attempts to switch models, but we do not need to follow this rigid pattern, for instance, we could randomly either update the current model or try to switch and we could vary the probabilities of updates and attempted model changes to try to improve overall mixing.
Here is a program that updates the current model 3/4 of the time and attempts a move between models 1/4 of the time.
program myRJMCMC
args logpost b ipar
local M = `b'[1,1]
if( runiform() < 0.75 ) {
* update within the current model
if( `M’ == 1 ) {
matrix `b'[1,2] = rbeta(13,17)
matrix `b'[1,3] = rbeta(26,24)
}
else {
matrix `b'[1,2] = rbeta(44,46)
matrix `b'[1,3] = rbeta(1,1)
}
}
else {
* try to switch
local psi1 = `b'[1,2]
local psi2 = `b'[1,3]
local a1 = log(binomialp(20,8,`psi1′))+log(binomialp(30,16,`psi2′))+ ///
log(betaden(5,5,`psi1′))+log(betaden(10,10,`psi2′))+log(0.3)
local a2 = log(binomialp(20,8,`psi1′))+log(binomialp(30,16,`psi1′))+ ///
log(betaden(20,20,`psi1′))+log(betaden(1,1,`psi2′))+log(0.7)
if `M’ == 1 {
if log(runiform()) < `a2′ – `a1′ matrix `b'[1,1] = 2
}
else {
if log(runiform()) < `a1′ – `a2′ matrix `b'[1,1] = 1
}
}
end
Once again the results are so similar that it is not worth showing them. The only noticeable difference is that the mixing is worse. This time we make around 400 model switches every 5000 iterations instead of the nearly 1800 that we achieved with the first program. As a result the estimate of P(M2|Y) is not as good and we ought to run the chain for longer.
Perhaps the main conclusion is, this is such a simple problem that virtually everything that we try will work perfectly well.
There is one further complication to put into the algorithm and that is to introduce a parameter transformation that will require Jacobians that are not equal to 1. Since this posting is already quite long and because the Jacobian is the element that is most likely to cause an error, I will cover this next time.


 Subscribe to John's posts
Subscribe to John's posts
Recent Comments